This is a second post on risk and effort in scientific research, prompted by this article. You can read the first post here.
It’s one of our first hikes in Missouri. For my husband and me, recently immigrated from southern California, everything seems just a little bit foreign. The spicy smell of sassafras, yellow midwestern light filtering through the leaves, the unbelievably deep blue sky. Our companions, two ecologists from my new department, teach us about glades—small rocky clearings that harbor different plants, insects, and animals than the surrounding forest (like prickly pear cactus!). We are headed to Taum Sauk, the highest peak in Missouri. We try not to snicker at the plaque stating that Taum Sauk is 1772 ft. high. (I mentally place air quotes around the word “peak”). But near the “top” we step off the shaded path onto a glade that provides as commanding a view as anyone could wish for, seemingly endless green hills fading to blue in the distance.
We aren’t just talking about the ecology of Missouri and making fun of its topology; we are also participating in a little solvitur ambulando. As we hike, I describe the fear and anxiety I’m feeling about getting a new project off the ground. “I’m so pumped about this idea, but what if we can’t get it to work? What if we have to punt in like 4 years, and then have nothing to show for it?” J. stops and turns around, his expression incredulous. “Nothing to show for it? I don’t even know what you mean! You have a hypothesis, you figure out how to test it, and then you report the results. If your hypothesis is wrong, you still learned something, right?”
Tool development is especially risky
I was equally incredulous to hear that some scientists’ work really does follow the classic hypothetico-deductive method of scientific inquiry. This had never been my experience. Almost every project I’d done as a plant or yeast molecular biologist carried substantial risk of not working, and not in an informative way. Usually, the risk came from the production of a tool. A tool could be a new DNA construct or a genetically modified model organism or an assay—something that needed to be in place before an interesting question could even be asked.
Once I was running my own lab, I still gambled on tool development, but now I depended on my lab members’ energy and time. In some cases we were successful, but—of course!—not always. My experience of one of the projects where I lost the tool development gamble still stings all these years later—both because I am really proud of the idea behind the project, and because am decidedly not proud of the way I executed it.
A cautionary tale
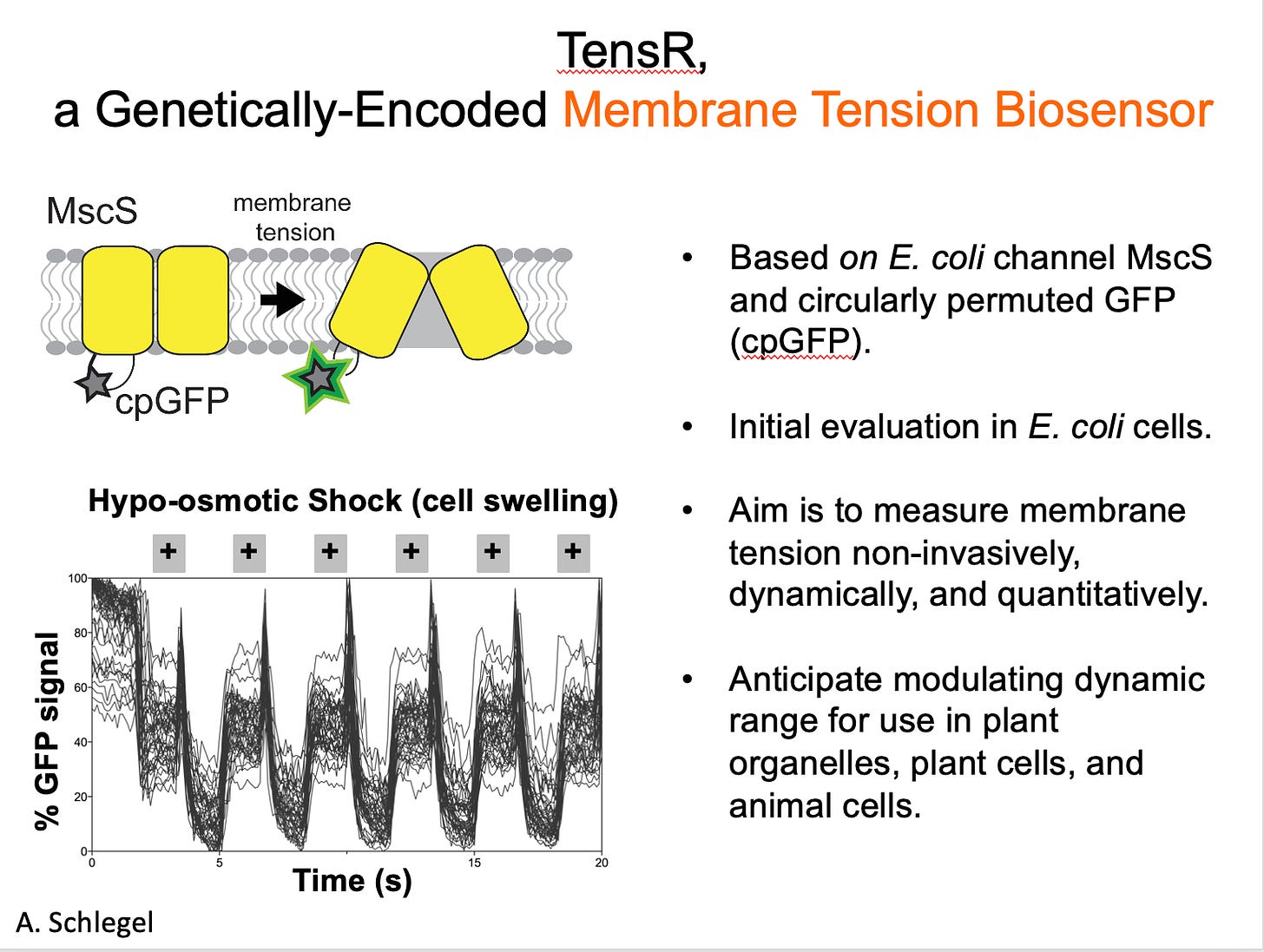
The concept was, I still think, super cool. We wanted to make a mechanical sensor, to turn a mechanosensitive ion channel into a molecular tool that could report on membrane tension. It was potentially exciting enough to be funded by multiple grants, and I was able to recruit a graduate student who agreed to take the project on. They built constructs, assessed the expression and stability of candidate sensors, and developed a methodology for measuring sensor function when expressed in bacteria. It was a classic high risk-high reward project.
At first, things looked great. We could see signal changing inside a bacterial cell whenever we changed its osmotic conditions, which should affect membrane tension. BUT, after months of experimental controls to test the tool, it wasn’t behaving as expected if it was responding to membrane tension, and the student did a final “kill” experiment that showed it wasn’t at all working the way we thought. And so the tool we’d been trying to build could not be built, at least not this particular way.
I also attempted to develop a genetically-encoded, fluorescence-based membrane tension sensor using the Mechanosensitive ion channel of Small conductance (MscS) from E. coli as a tension-sensing scaffold and circularly permuted GFP (mcpGFP) as the fluorescence reporter. While responses to tension by sensor candidates could not be ruled out, signal changes from mcpGFP that were not dependent on the tension-sensing scaffold dominated sensor responses to osmotic shock.—A. Schlegel, PhD thesis
We discussed, extensively, what to do next. Should we try to figure out what our candidate sensor WAS responding to? Should we test out a different mechanosensitive ion channel, or choose a different assay? Should we write up what we’d observed so far as a cautionary tale?
Risk and Regret
But in the end we did none of these things. The graduate student working on the project was eager to put their focus onto a more straightforward project they’d been moving along in the background, and I wasn’t able to convince anyone else to sink time or effort into a project that had already proven so challenging. Those data didn’t make it into a peer-reviewed publication, and now that I’ve left academia, they never will. And I do regret that, a little. Surely plenty of others had a similar idea and would like to know what NOT to do. To return to the ecologist’s point back then in the Missouri forest—”you still learned something, right?”—yes, we did learn something and in an ideal world we would have published it. Certainly that would have been the right way to serve the foundations that supported the research.
But what I regret far more is how this project impacted the graduate student who worked on it. They gained knowledge and expertise—it was definitely a mechanism for training. But it was too much to take on: a project with high potential, in which we had almost no lab expertise, for which so many components had to be developed, and where we knew we had a lot of competition. I didn’t do enough to protect them from the pressure of it all. For me, the project was fun to talk about and the promising preliminary data helped me get other grants. For them, I suspect that the stress and repeated disappointment re-shaped their relationship to bench science, taking the pleasure and fun and excitement out of it.
I guess what I’m trying to say here is that risk is relative. I know now: lab leaders should think about and discuss with their trainees how risky projects might affect their CVs, graduation time, and their overall experience and motivation. What one person might consider a laughable walk up a hill, another might experience as a hike to the highest point in the state.
Discussion Section
Have you had an experience working on a risky project? How did it affect your career? Did it change your relationship with science?



Ugh, this hits me hard! Working on a research organism that isn't one of the established ones, I have always had to do tool development myself, I actually enjoy the tinkering and wish we could have more freedom to do that. I have been trying to balance things for my trainees by getting them started on both risky and bread-and-butter projects. One way I proactively help them reframe the "fails" is that "success is measured by whether you are trying new things and troubleshooting, not by positive results". But, I also tell them, we need papers, and we should check in often about the progress to decide whether we should move onto something else. It's always an ongoing reassess and adjust.
I wonder if preprint servers like bioRXiv, now used so much more commonly, can help to bridge that gap. Writing up a pile of rigorously performed experiments but negative data in anticipation of peer reviewers inevitably asking for yet more experiments is enough to make even the most enthusiastic trainee (or PI) throw up their hands. Getting this info out as a a preprint at least gets it into the public record, although it doesn't quite as well address your main point of risk to the trainee. Originally it seemed like the idea of journals like PLoS One or some of the solid society journals was that people could publish stories that were technically rigorous but didn't necessarily come to a flashy conclusion. In practice, even for these journals, reviewers still sometimes ask for more experiments to expand the scope rather just assessing the rigor of the experiments presented.